Stefano Vanni
SNSF Professor, ERC investigator
PER 05 - 1.301
+41 26 300 8896
E-mail
Our main goal is to understand how the molecular properties of proteins and membranes modulate cellular processes. To do so, we develop new computational approaches to investigate soft matter in silico, and we combine these investigations with biochemical and biophysical approaches. Currently, the main focus of the lab is to understand how specific lipids and membrane properties influence intracellular trafficking processes and fat storage in eukaryotic cells.
Our research focuses on the following topics
Our main approach is molecular dynamics (MD) simulations. Using this approach, we can describe molecular systems in the range of 1-100 nanometers with atom-level accuracy. To use Feynman words, we investigate living matter by studying the “the jiggling and wiggling of atoms”. To do so, we are currently developing new force-field parameters to investigate large-scale cellular phenomena involving membranes with unprecedented chemical accuracy using MD approaches.

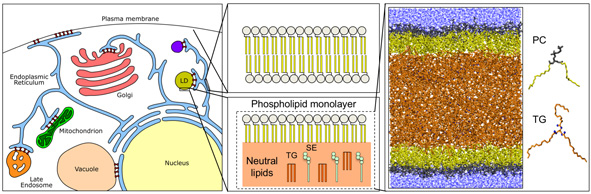
Lipids are characterized by their amphipathic character. Because of this, they typically arrange in a bilayer configuration. The hydrophobic tails, however, also serve as a reservoir of metabolic energy, undergoing beta-oxidation in the Krebs cycle. In cells, lipids are stored in specialized organelles called lipid droplets (LDs) after esterification of their polar head with an additional hydrophobic chain. Thus, LDs can be considered as the dispersed phase of oil-emulsions-in-water, with the cytosol as the continuous phase. Our understanding of the molecular properties of LDs is very limited, and we are currently investigating how their function and behaviour is modulated by their unique chemical properties.
Cell membranes are continually remodelled to achieve communication between intracellular compartments and to selectively exchange materials between them. The energetics of these remodelling processes are governed by the interplay between specialized proteins and membrane properties, but in most cases, we still lack a detailed molecular explanation of how these processes can be modulated in a cellular environment. By combining coarse grain MD simulations and biochemical and cellular experiments, we investigate how membrane properties modulate remodelling processes and how this might influence cell functioning.

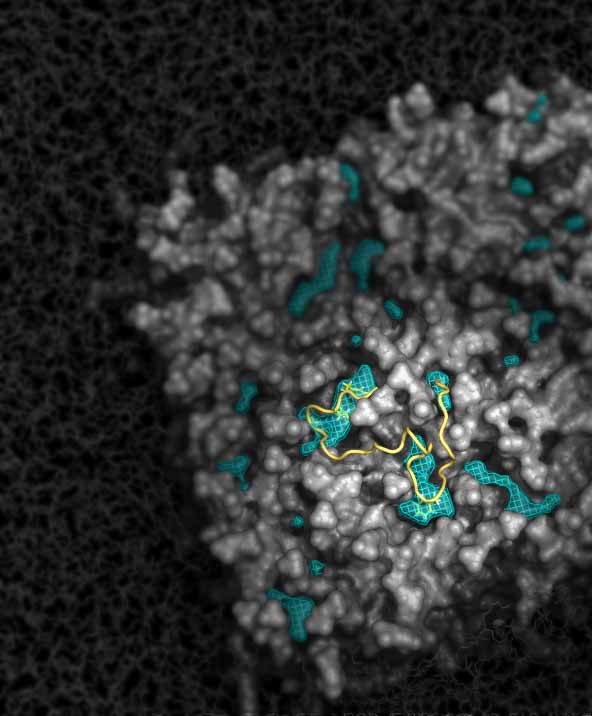
Interactions between proteins and membranes are a vital part of cell functioning. Some proteins only interact with membranes transiently, while others are constantly embedded into the membrane. In both cases, however, proteins have been engineered by nature to “sense” their membrane environment and to respond to its variations. Using a combination of MD simulations and biochemical approaches, we study how membrane properties regulate protein functions, and hence how lipid metabolism may play a role in unexpected, and apparently unrelated, trafficking processes.

SNSF Professor, ERC investigator
PER 05 - 1.301
+41 26 300 8896
E-mail